-
In the last 10 years we have witnessed tremendous progress in breast cancer management. Thanks to the advances in the field, female breast cancer death rate has significantly dropped, which is quite impressive when considering the fact that the rate had been continually increasing since 1980[1]. Early detection by mammography and hormone replacement therapy have significantly reduced death rate of female breast cancer patients and have greatly improved their quality of life[2-3]. However, as complexity and variety within breast cancers have been increasingly recognized in recent years, tailored management strategies that are specific to various subtypes of breast cancers can hold great promises in further improving our current treatment regimens of this important disease. Thus, characterization of breast cancers at the molecular level has become the center of focus in breast cancer research.
Of all these molecular biomarkers, human epidermal growth factor receptor(EGFR) family plays a central role in breast cancer initiation and progression. In EGFR family, the importance of human epidermal growth factor receptor 2(HER2) has been extensively studied as a predictive marker in breast cancer and HER2 targeted therapy has achieved much success[4-6]. However, only about 25%-30% of breast cancer has been identified as HER2 positive tumor. Triple-negative breast cancer (TNBC: ER-/PR-/HER2-) represents 15% of newly diagnosed breast cancer and is overrepresented in younger patient population and patients with African ancestry[7]. As for diagnosis and treatment, TNBC is almost always missed by mammography and have very poor prognosis even with standard-of-care chemotherapy. Therefore it is of heightened importance to establish an effective molecular biomarker for specific and effective therapy for patients suffering from this highly malignant subtype of breast cancer.
Anti-EGFR monoclonal antibody Cetuximab has been approved for treatment of EGFR-positive metastatic colorectal cancer since 2004 and for treatment of advanced squamous cell carcinoma of the head and neck in combination with radiation in 2005[8-9]. More than 50% of TNBC overexpress EGFR and may represent a group of breast carcinoma that could benefit from EGFR targeted therapy[10-11]. Studies have shown EGFR as a negative prognostic factor in TNBC patients[12-14]. The combination of tyrosine kinase inhibitor Gefitinib with conventional chemotherapy has shown improved efficacy compared with chemotherapy or targeted therapy alone in TNBC[10]. The combination of Cetuximab and cisplatin showed efficacy in MDA-MB-468 breast carcinoma cell[15]. Triple combination of Gefitinib, carboplatin and docetaxel has been proved to be synergistic in TNBC cells[10]. However, a lack of standardization in approaches to definition of EGFR dysregulation leads to uncertainties in establishing EGFR as a prognostic and predictive factor in breast cancer, particularly in vivo. We believe that the role of EGFR in TNBC could be further investigated by comprehensive in vivo profiles of EGFR expression monitored by noninvasive imaging approaches[16]. More importantly, quantitative PET studies of EGFR expression profile could provide a screening process and subsequently direct targeted therapy towards EGFR as well as predict its prognosis. Herein, we propose to use PET for in vivo characterization of EGFR expression in TNBC.
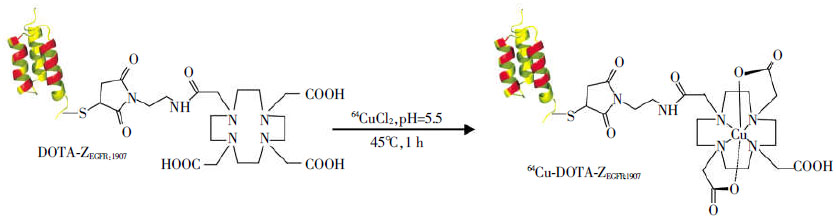
The development of molecular probes provides diverse platforms for imaging molecular events associated with different cancers. Of these platforms, antibody and antibody fragments have been around for more than 50 years and have gained wide popularity[17]. However, the slow in vivo pharmacokinetics of intact antibody and suboptimal tumor uptake of smaller fragments of antibody may limit their use as ideal probes, especially when they are tagged with short-half radionuclides such as 18F. Recently non-immunogenic scaffold protein Affibody molecules have drawn a lot of attention for developing affinity ligands against a variety of molecular targets[18-19]. Affibody molecules were derived from one of the Immunoglobulin G(IgG)-binding domains of staphylococcal protein A, and they are composed of a relatively small engineered non-immunoglobin protein scaffold with 58 amino acid residues and a 3-α-helical bundle scaffold structure. Affibody molecule libraries can be easily constructed by randomization of 13 amino acid residues in helices 1 and 2 of the 3-helix bundle protein. Thus Affibody binders with high affinities and specificities against a wide variety of desired targets can be quickly identified and selected using phage-display libraries technology and affinity maturation[20-23]. Anti-EGFR affibody has been developed to selectively bind to extracellular domain of EGFR with nanomolar affinity[24-25]. It can be used to target various tumors with overexpression of EGFR and has shown promising results[26-28]. Besides that an promising imaging probe, anti-EGFR affibody could activate Erk and Akt without detectable EGFR autophosphorylation, thus could be used for EGFR targeted therapy in cancer[29]. In our previous studies, an anti-EGFR affibody analogue, Ac-Cys-ZEGFR:1907, was chemically synthesized using a solid phase peptide synthesizer and then site-specifically conjugated with 1, 4, 7, 10-tetraazacyclododecane-1, 4, 7, 10-tetraacetic acid (DOTA) for radiolabeling with 64Cu or N-2-(4-18F-fluorobenzamido)-ethyl]-maleimide (18F-FBEM) to successfully prepare 64Cu-DOTA-ZEGFR:1907 or 18F-FBEM- ZEGFR:1907, respectively[30-31](Fig. 1). In this study, we further studies whether 64Cu-DOTA-ZEGFR:1907 can serve as a PET probe for in vivo imaging of EGFR expression in TNBC small animal models.

Figure 1. Western blot analysis of epidermal growth factor receptor expression in different breast cancer cell lines (duplicate samples per cell line)(A). Cell uptake of 64Cu-DOTA-ZEGFR:1907 in MCF7 and SUM159 cells over time at 37℃with or without the presence of non-radioactive affibody molecules ZEGFR:1907 (B). All results were expressed as mean of triplicate measurement ±standard deviation.
-
Maleimido-mono-amide-DOTA was obtained from Macrocyclics Inc.(Dallas, TX). (S)-2-(Fmoc-amino)-4-tritylsulfanyl-butyric acid(Fmoc-HomoCys(Trt)-OH) was purchased from Bachem Bioscience, Inc(King of Prussia, PA). All other N-α-Fmoc-protected amino acids were purchased from Advanced Chemtech (Louisville, KY). Dimethylformamide(DMF) and methylene chloride were from Fisher Scientific(Fair Lawn, NJ). Piperidine(20%) in DMF and 0.4 mol/L of N-methylmorpholine(NMM) in DMF were from Protein Technologies Inc.(Tucson, AZ). Trifluoroacetic acid(TFA), O-benzotriazole-N, N, N′, N′-tetramethyluronium hexafluorophosphate(HBTU), and 4-(2′, 4′-dimethoxy-phenyl-Fmoc-aminomethyl)-phenoxy resin(Rink amide resin LS, 100-200 mesh, 1% DVB, 0.2 mmol/g) were from Advanced Chemtech. Pyridine, acetic anhydride, acetic acid, and anhydrous ether were from J.T.Baker (Phillips-burg, NJ). Triisopropylsilane(TIPS) and 1, 2-ethanedi-thiol(EDT) were purchased from Aldrich (Milwaukee, WI). High performance liquid chromatography(HPLC) grade acetonitrile(CH3CN) and Millipore 18 mΩ water were used for peptide purifications. All other standard synthesis reagents were purchased from Sigma-Aldrich Chemical Co.(St. Louis, MO). The radionuclide 64Cu was provided by the Department of Medical Physics, University of Wisconsin at Madison(Madison, WI). All the other general materials(cell lines, mice, etc.) and instruments(reverse phase HPLC, radioactive dose calibrator, Mass spectrometer) are the same as that previously reported[30].
-
Ac-Cys-ZEGFR:1907(Ac-CVDNKFNKEMWAAWEEI-RNLPNLNGWQMTAFIASLV DDPSQSANLLAEAKK-LNDAQAPK-NH2) was synthesized on an automatic peptide synthesizer(CS Bio, CS 336X) as described in our previous reports[30-31]. Briefly, peptide was synthesized using standard solid phase peptide synthesizer (Fmoc chemistry) and then purified by a reversed phase preparative HPLC with a protein and peptide C4 column(Vydac, Hesperia, CA). The identity of the target peptides was confirmed by matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF-MS, model: Perseptive Voyager-DE RP Biospectrometer) (Framingham, MA) or an electrospray ionization time of flight mass spectrometer(ESI-TOF-MS, model: JMS-T100LC) (JEOL, Tokyo, Japan).
-
The DOTA-ZEGFR:1907 was radiolabeled with 64Cu as previously described[30]. Briefly, about 111 MBq 64CuCl2 was added into 20 μg DOTA-ZEGFR:1907 in 0.1 N NaOAc(pH 5.5) buffer followed by a 60 min incubation at 45 ℃. EDTA(5 μL, 10 mmol/L) was then added to quench the free 64Cu. The radiolabeled complex was further purified by a PD-10 column(GE Healthcare, Piscataway, NJ). The product was washed out by phosphate-buffered saline(PBS) and passed through a 0.22 μm Millipore filter into a sterile vial for in vitro and animal experiments. Radioanalytical HPLC was used to analyze the purified 64Cu-DOTA-ZEGFR:1907. The flow rate was 1 mL/min, with the mobile phase starting from 95% solvent A(H2O/0.1%TFA) and 5% solvent B(CH3CN/0.1%TFA) (0-3 min) to 35% solvent A and 65% solvent B at 33 min, then going to 15% solvent A and 85% solvent B(33-36 min), maintaining this solvent composition for another 3 min(36-39 min) and returning to initial solvent composition by 42 min.
-
TNBC SUM159 cells were cultured in Dulbecco′s Modified Eagle Medium (DMEM)/F15 medium (Mediatech, VA) supplemented with 10% fetal bovine serum (FBS), 1% penicillin-streptomycin (Invitrogen Life Technologies, Carlsbad, CA), insulin and hydrocortisone. TNBC MDA-MB-231 cells were maintained in DMEM supplemented with 4 mmol/L L-Glutamine, 10% FBS, and 1% penicillin-streptomycin. ER positive MCF7 breast cancer cells were cultured in DMEM (high glucose), supplemented with 10% fetal bovine serum(FBS), 0.1 mmol/L MEM Non-Essential Amino Acids(NEAA), 2 mmol/L L-glutamine, and 1% penicillin-streptomycin. All these cells were maintained in a humidified atmosphere of 5% CO2 at 37℃, with the medium changed every other day. A 70%-80% confluent monolayer was detached by 0.1% trypsin and dissociated into a single cell suspension for further cell culture.
MDA-MB-231, SUM149, and MCF7 cell lysates was extracted using RIPA tissue protein extraction buffer and the concentration was determined using microBCA protein assay kit (Pierce Biotechnology, Inc., Rockford, IL). After sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) separation of 30 μg of total protein, protein was transferred to a polyvinylidene fluoride membrane (Invitrogen Corp., Carlsbad, CA) and incubated at room temperature with 5% non-fat milk blocking buffer. The blots were then incubated overnight at 4℃ with anti-EGFR antibody (Cell signaling technology), followed by incubation at room temperature for 1 h with HRP-conjugated anti-human antibody(GE Healthcare, Piscataway, NJ). The bands were detected using the ECL western blotting reagent(GE Healthcare), and imaged using the Xenogen IVIS 100 imaging system(exposure time 1 min; binning 4; no filter; f/stop 1). Actin was used as a loading control.
-
The in vitro cell uptake assays of the 64Cu-DOTA-ZEGFR:1907 were performed with the SUM159 and MCF7 cells. Briefly, about 106 cells were harvested and resuspended with serum free medium containing 64Cu-DOTA-ZEGFR:1907 (37 kBq). The cells were incubated for 1 h and 2 h under 37℃. The nonspecific binding of the probes with SUM159 cells were then determined by co-incubation with non-radiolabeled ZEGFR:1907 molecules [1 μg of ZEGFR:1907 was added to each sample]. The cells were rinsed three times with cold PBS (0.01 mol/L, pH=7.4)/0.2% BSA and the radioactivity of the cell pellets were counted using PerkinElmer 1470 automatic gamma counter. The cell uptake was determined by radioactivity associated with cells divided by the total radioactivity added and then time 100.
-
All animal studies were carried out in compliance with federal and local institutional rules for the conduct of animal experimentation. MDA-MB-231 and SUM159 cells were harvested and 3×106 cells were diluted in 100 μL of PBS then subcutaneously implanted into the right upper shoulder of 6-7-week-old female nu/nu mouse(Charles River, Wilmington, MA). Tumors were allowed to grow to a size of 0.5-1.0 cm in diameter(3-6 weeks), and the tumor bearing mice were subjected to in vivo biodistribution and imaging studies.
-
All mice bearing SUM159 tumors(n=4 for each group) were injected with 64Cu-DOTA-ZEGFR:1907 (1.92-2.07 MBq, 1.3-1.4 μg) through the tail vein and sacrificed at 1, 4 and 24 h post-injection(p.i.). Tumor and normal tissues of interest were excised and weighed, and their radioactivity was measured in a Wallac 1480 automated-counter(Perkin Elmer, MA). The radioactivity uptake in the tumor and normal tissues was expressed as a percentage of the injected radioactive dose per gram of tissue(% ID/g).
-
PET imaging of tumor-bearing mice(MDA-MB-231 or SUM159) was performed on a small animal PET R4 rodent model scanner(Concord Microsystem, Knoxville, TN) and Inveon PET-CT(Siemens). The mice bearing MDA-MB-231 or SUM159 tumors(n=4 per group) were injected with 64Cu-DOTA-ZEGFR:1907(2 MBq, 1.4 μg) through lateral tail vein and imaged at 1, 4, 24 and 48 h post-injection(p.i.) of the probe. For another group of mice bearing SUM159 tumors(n=4 per group), each mouse was co-injected with the same activity of probe and 25 or 125 μg of non-radiolabeled ZEGFR:1907. At different times p.i.(1, 4 and 24 h), the mice were anesthetized with 2% isofluorane, and placed in the prone position and the center of the filed of view of microPET. The 5 min static scans were obtained and the images were reconstructed by a two-dimensional ordered subsets expectation maximum (OSEM) algorithm.
-
Statistical analysis was performed using the Student′s t-test for unpaired data. A 95% confidence level was chosen to determine the significance between groups, with P < 0.05 being significantly different. All statistical computations were processed by the Microsoft Office Excel 2007(Microsoft, Redmond, VA).
-
Ac-Cys-ZEGFR:1907 with a cysteine residue at the N-terminal was successfully synthesized through conventional solid phase peptide synthesis and purified by semi-preparative HPLC. The measured molecular weight(MW) was consistent with the expected MW: m/z=6690.0 for[M+H]+(calculated MW[M+H]+=6689.6). The purified peptide was generally obtained in about 10% yield. Mass spectrometry analysis of the final product also only showed the expected mass peak for DOTA-ZEGFR:1907. The measured MW was m/z=7215.0 for[M+H]+ (calculated MW[M+H]+=7215.1). The recovery yield was over 40% after purification, and the purity for the final product was > 95%(retention time: 26 min). The DOTA-ZEGFR:1907 peptide was then radiolabeled with 64Cu. The purification of radiolabeling solution, using a PD-10 column, afforded 64Cu-ZEGFR:1907 with > 95% radiochemical purity with modest specific activity 4.25-8.5 MBq/nmol. HPLC analysis showed its retention time was ~26 min. Serum stability of 64Cu-DOTA-ZEGFR:1907 has been previously shown to be excellent with > 95% of the probes intact after 4 h incubation and approximately 85% of the probes intact after 24 h incubation[30].
-
Western blot assays showed that MDA-MB-231 had the highest EGFR expression among three cells tested, and SUM159 displayed moderate EGFR expression, whereas MCF7 exhibited low EGFR expression(Fig. 2A). Then the SUM159 cells with moderate EGFR expressing and MCF7 with low expression were used for evaluation of the in vitro EGFR binding ability and specificity of 64Cu-DOTA-ZEGFR:1907. Cell uptake of the probe at 37℃ over a 2 h incubation period was shown in Fig. 2B. At 1 and 2 h, the uptake of the probe in SUM159 cells showed (3.92±0.24)% and (2.74±0.31)% of applied radioactivity, respectively. While only (0.87±0.11)% and (0.47±0.10)% was observed at these two points, when the cells were incubated with the probe and large excess of affibody molecules ZEGFR:1907. This significant(t=25.3, 14.7, both P < 0.05) inhibition of the probe uptake clearly indicated the EGFR binding specificity of the probe. Moreover, the MCF7 cell only showed (1.63±0.41)% and (1.59±0.17)% of applied radioactivity at 1 and 2 h, respectively, further indicating the in vitro targeting specificity of the probe.

Figure 2. Western blot analysis of epidermal growth factor receptor expression in different breast cancer cell lines (duplicate samples per cell line)(A). Cell uptake of 64Cu-DOTA-ZEGFR:1907 in MCF7 and SUM159 cells over time at 37℃ with or without the presence of non-radioactive affibody molecules ZEGFR:1907(B). All results were expressed as mean of triplicate measurement ± standard deviation.
-
Biodistribution results for 64Cu-DOTA-ZEGFR:1907 at 1, 4 and 24 h p.i. were summarized in Table 1. SUM159 tumor uptake gradually increased from (1.6± 0.3) %ID/g at 1 h to(4.1±0.9) %ID/g at 24 h. The blood uptake decreased from(6.3±1.2) %ID/g at 1 h p.i. to (2.2±1.2) %ID/g at 24 h p.i. Very high renal uptake and was found for the probe with (73.4±1.6) %ID/g at 4 h and reached 85.3±4.2 at 24 h p.i., which matched with previous finding when using radiometal labeled affibody as the targeted molecules[30, 32-35]. The liver uptake was (15.8±4.6) and (13.0±2.1) %ID/g at 1 and 24 h p.i., respectively. Most of other normal organs showed low uptake of the probe(Table 1, Fig. 3A). The clearance of 64Cu-DOTA-ZEGFR:1907 from the blood caused the tumor to blood ratio reached 2.2 at 24 h. The tumor to muscle ratio could reach 5.7 at 24 h p.i.(Table 1, Fig. 3B).

Figure 3. Biodistribution results(A) and tumor to normal tissue ratios(B) for 64Cu-DOTA-ZEGFR:1907 in nude mice bearing subcutaneously xenotransplanted SUM159 human breast cancer
Organ 1 h 4 h 24 h Tumor 1.6±0.3 3.3±1.7 4.1±0.9 Blood 6.3±1.2 5.9±1.6 2.2±1.2 Heart 1.9±0.6 2.0±0.5 2.1±0.5 Liver 15.8±4.6 20.4±4.8 13.0±2.1 Lung 2.9±0.5 2.8±0.2 3.1±0.6 Muscle 0.3±0.1 0.9±0.4 0.8±0.2 Spleen 1.3±0.3 1.5±0.3 2.1±0.2 Brain 0.2±0.0 0.3±0.1 0.3±0.1 Intestine 2.0±0.4 1.9±0.2 3.3±0.6 Skin 1.1±0.3 2.6±0.4 2.1±0.9 Stomach 1.5±0.8 2.0±0.4 2.6±0.8 Pancreas 1.7±0.7 2.0±0.3 1.6±0.6 Bone 0.8±0.3 0.6±0.3 0.7±0.3 Kidney 35.6±12.7 73.4±1.6 85.3±4.2 Tumor/normal organ ratio Tumor/Blood 0.25±0.07 0.65±0.51 2.22±1.13 Tumor/Muscle 5.21±1.35 4.52±3.14 5.70±1.98 Table 1. Biodistribution data for 64Cu-DOTA-ZEGFR:1907 in nude mice bearing subcutaneously xenotransplanted SUM159 human triple negative breast cancer. Data are expressed as the percentage administered activity(injected dose) per gram of tissue(%ID/g) after intravenous injection of 0.69-1.01 MBq probe at 1, 4 and 24 h p.i.(n=4).
-
Representative decay-corrected coronal small animal PET images of nude mice bearing MDA-MB-231 (high EGFR expression) and SUM159 (moderate EGFR expression) tumors on their right shoulder at 1, 4, 24 and 48 h p.i. are shown in Fig. 4. Both tumors were clearly delineated with good tumor to contralateral background contrast at 4, 24 and 48 h p.i.. Moreover, the MDA-MB-231 displayed better imaging contrast than that of SUM159 at 4 and 24 h p.i..
Decay-corrected coronal small animal PET images of a mouse bearing SUM159 injected with the probe and unlabeled ZEGFR:1907(25 μg top row; or 125 μg bottom row) at 1, 4 and 24 h are shown in Fig. 5. For 64Cu-DOTA-ZEGFR:190 co-injected with 25 μg cold affibody Ac-CysZEGFR:1907, SUM159 tumor was clearly visualized with high tumor-to-background contrast from 1 to 24 h p.i. Liver activity was observable while high activity accumulation was particularly obvious in the kidney region, which was consistent with the findings from the biodistribution study. Furthermore, the co-injection of unlabeled Ac-Cys-ZEGFR:1907 125 μg significantly reduced the uptake of 64Cu-DOTA-ZEGFR:1907 in the tumor and liver, resulting in much lower target to background contrast in vivo(Fig. 5). Tumor uptake after co-injection of 25 or 125 μg of unlabeled ZEGFR:1907 and sacrificed at 4 h p.i. Further quantification analysis indicated that tumor uptake was blocked by 48.2%.
-
A variety of radiolabeled Affibody molecules have been successfully demonstrated to be promising molecular probes for imaging tumor biomarkers such as EGFR expression in living mice[30-35]. Particularly, highly clinically relevant 18F and 64Cu-labeled EGFR Affibody have also been evaluated in tumor bearing mice by our group[30-31]. In this study, the potential advantages expected for smaller protein constructs including, economic viability, potentially low immunogenic potential, high specific tumor targeting ability, fast tumor extravasation, and reasonable tumor accumulation and retention have been again shown in a clinically relevant scenario: PET imaging of triple negative breast cancer. With many reports on high EGFR expression in TNBC, we are very keen to evaluate the EGFR protein expression in TNBC models and to explore the opportunities of using EGFR as an in vivo theranostic marker for TNBC. This study provides us with first-hand profiles of EGFR expression patterns as well as the in vivo behaviors of 64Cu-DOTA-ZEGFR:1907 in TNBC models.
The tumor targeting ability and pharmacokinetics of 64Cu-DOTA-ZEGFR:1907 was first evaluated in a TNBC model, mice bearing SUM159 tumor. The probe displayed good tumor uptakes at 4 h p.i. and 24 h in SUM159 models(Table 1). Reasonable tumor-to-blood and tumor-muscle ratios(2.22 and 5.70) were attained at 24 h p.i., suggesting 64Cu-DOTA-ZEGFR:1907 can be used for tumor imaging. Lower tumor-to-blood ratio (0.65) was achieved at 4 h, mainly because of the relatively high activity residue in the blood. But good tumor-to-muscle ratio(4.52) at 4 h still warrants the good tumor imaging quality(Fig. 4 and 5). High uptake in kidney and moderate liver and intestines uptakes were also observed, indicating that the probe was mainly cleared through kidney-urinary system and partially cleared through hepatobiliary system. Many reports have revealed that radiometal labeled affibody molecules displayed high kidney uptake and retention, which may limit their applications for tumor imaging and treatment, especially when affibody is labeled with therapeutic radionuclides[32-35].
The in vivo tumor imaging ability of 64Cu-DOTA-ZEGFR:1907 was further examined using mice bearing TNBC MDA-MB-231 or SUM159 with high to medium EGFR expression and small animal PET imaging. The EGFR expression pattern has been explored in many breast cancer cell lines. We are the first group to use affibody based PET imaging approaches to evaluate EGFR expression in TNBC. Our study clearly demonstrates that 64Cu-DOTA-ZEGFR:1907 can successfully image TNBCs with high and moderate EGFR expression (Fig. 4). Excellent tumor to background contrast can be obtained as early as 1 h after intravenous injection of the probe(Fig. 5), confirming the fast tumor targeting and clearance of the affibody molecule. Good tumor localization and contrast have also been observed at 24 h p.i.; consistent with the finding obtained in biodistribution study. Similarly, SUM159 tumors were not visible in the mice with co-injection of excess of large amount of cold affibody molecules, suggesting good specificity of the probe in vivo(Fig. 5).
In summary, an Affibody based EGFR imaging approaches has been successfully established to monitor EGFR profile in triple negative breast cancer models. Radiolabeled ZEGFR:1907 small protein shows good in vitro and in vivo tumor targeting ability. Biodistribution and small animal PET imaging studies further demonstrate that 64Cu-labeled EGFR affibody is a promising molecular probe for imaging EGFR positive tumor in TNBC. Strategies to further optimize the biodistribution of the ZEGFR:1907 are under active investigation in our laboratories.
Conflict of interest The authors declare no conflicts of interest.
Authors contribution statement Yingding Xu wrote the manuscript in addition to designing, performing, and analyzing all experiments. Gang Ren designed, performed, and analyzed all experiments. Shibo Qi synthesized and characterized the probe 64Cu-DOTA-ZEGFR:1907. Zhen Cheng designed, supervised, and analyzed all experiments, in addition to assisting with manuscript preparation.
Profiling EGFR in Triple Negative Breast Tumor Using Affibody PET Imaging
Profiling EGFR in Triple Negative Breast Tumor Using Affibody PET Imaging
-
Abstract: Objective Triple negative breast cancer(TNBC) represents a group of refractory breast cancers with aggressive clinical manifestations as well as poor prognoses. Human epidermal growth factor receptor(EGFR) expression is strongly associated with TNBC progression and it may serve as a therapeutic target for TNBC. We aimed to evaluate EGFR affibody-based PET imaging to profile EGFR expression in small animal models. Methods 1,4,7,10-tetraazacyclododecane-1,4,7,10-tetraacetic acid(DOTA) conjugated Ac-Cys-ZEGFR:1907 was chemically synthesized using solid phase peptide synthesizer and then radiolabeled with 64Cu. The in vitro cell uptake study was performed using SUM159 and MCF7 cells. The biodistribution and small animal PET imaging using 64Cu-DOTA-ZEGFR:1907 were further carried out with nude mice bearing subcutaneous MDA-MB-231 and SUM159 tumors. Results DOTA-Ac-Cys-ZEGFR:1907 was successfully synthesized and radiolabeled with 64Cu. Biodistribution study showed that tumor uptake value of 64Cu-DOTA-Ac-Cys-ZEGFR:1907 remained at(4.07?.93)%ID/g at 24 h in nude mice(n=4) bearing SUM159 xenografts. Furthermore, small animal PET imaging study clearly showed that 64Cu-DOTA-Ac-Cys-ZEGFR:1907 specifically delineated the EGFR positive TNBC tumors at 4 h or later. Conclusion The study demonstrates that 64Cu-DOTA-Ac-Cys-ZEGFR:1907 is a promising molecular probe for PET imaging of EGFR positive TNBC. EGFR based small protein scaffold holds great promise as a novel platform that can be used for EGFR profiling of TNBC.
-
Figure 1. Western blot analysis of epidermal growth factor receptor expression in different breast cancer cell lines (duplicate samples per cell line)(A). Cell uptake of 64Cu-DOTA-ZEGFR:1907 in MCF7 and SUM159 cells over time at 37℃with or without the presence of non-radioactive affibody molecules ZEGFR:1907 (B). All results were expressed as mean of triplicate measurement ±standard deviation.
Figure 2. Western blot analysis of epidermal growth factor receptor expression in different breast cancer cell lines (duplicate samples per cell line)(A). Cell uptake of 64Cu-DOTA-ZEGFR:1907 in MCF7 and SUM159 cells over time at 37℃ with or without the presence of non-radioactive affibody molecules ZEGFR:1907(B). All results were expressed as mean of triplicate measurement ± standard deviation.
Table 1. Biodistribution data for 64Cu-DOTA-ZEGFR:1907 in nude mice bearing subcutaneously xenotransplanted SUM159 human triple negative breast cancer. Data are expressed as the percentage administered activity(injected dose) per gram of tissue(%ID/g) after intravenous injection of 0.69-1.01 MBq probe at 1, 4 and 24 h p.i.(n=4).
Organ 1 h 4 h 24 h Tumor 1.6±0.3 3.3±1.7 4.1±0.9 Blood 6.3±1.2 5.9±1.6 2.2±1.2 Heart 1.9±0.6 2.0±0.5 2.1±0.5 Liver 15.8±4.6 20.4±4.8 13.0±2.1 Lung 2.9±0.5 2.8±0.2 3.1±0.6 Muscle 0.3±0.1 0.9±0.4 0.8±0.2 Spleen 1.3±0.3 1.5±0.3 2.1±0.2 Brain 0.2±0.0 0.3±0.1 0.3±0.1 Intestine 2.0±0.4 1.9±0.2 3.3±0.6 Skin 1.1±0.3 2.6±0.4 2.1±0.9 Stomach 1.5±0.8 2.0±0.4 2.6±0.8 Pancreas 1.7±0.7 2.0±0.3 1.6±0.6 Bone 0.8±0.3 0.6±0.3 0.7±0.3 Kidney 35.6±12.7 73.4±1.6 85.3±4.2 Tumor/normal organ ratio Tumor/Blood 0.25±0.07 0.65±0.51 2.22±1.13 Tumor/Muscle 5.21±1.35 4.52±3.14 5.70±1.98 -
[1] [2] [3] [4] [5] [6] [7] [8] [9] [10] [11] [12] [13] [14] [15] [16] [17] [18] [19] [20] [21] [22] [23] [24] [25] [26] [27] [28] [29] [30] [31] [32] [33] [34] [35] -




 下载:
下载: